SDSC/UCSD Study Uncovers Mechanisms of Cancer-Causing Mutations
Computer modeling leads to more precise targeting of therapies
Published Date
Article Content
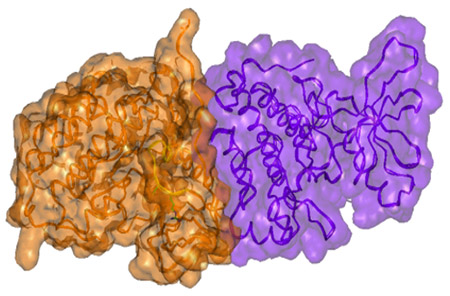
Surface presentation of the active dimer of the EGFR kinase subunits. Note the close complementary interaction of the subunits. Igor F. Tsigelny, SDSC/UC San Diego
Researchers at the San Diego Supercomputer Center (SDSC) and the Moores Cancer Center at the University of California, San Diego, have described for the first time the molecular mechanism of cancer development caused by well-known “resistance” mutations in the gene called epidermal growth factor receptor (EGFR).
While these mutations were known for quite a long time, the question as to why they cause cancer or make some drugs ineffective was still not answered.
The study, called “Molecular Determinants of Drug-Specific Sensitivity for Epidermal Growth Factor Receptor (EGFR) Exon 19 and 20 Mutants in Non-Small Cell Lung Cancer,” and published online in the journal Oncotarget, demonstrates how computer modeling of EGFR mutations found in lung cancer can elucidate their molecular mechanism of action and consequently optimize the selection of therapeutic agents to treat patients.
On the basis of structural analysis of this protein, scientists demonstrated that specific mutations within the EGFR gene increase electrostatic interactions between the two dimer subunits of EGFR, stabilizing it in an “active” state, leading to cancer. Such a finding suggested that the treatment of EGFR-related lung cancers with the usual small molecule inhibitory drugs would not be successful (patients harboring these mutations are typically resistant to these drugs), while antibodies that interfere with dimerization of EGFR would be effective.
“The mutations are substitutions of one amino acid to another, or a set of neighboring amino acids to a different set of them,” said lead author Igor F. Tsigelny, a research scientist with the SDSC as well as the UC San Diego Moores Cancer Center and the Department of Neurosciences. “The proteins have mechanisms of adaptations for possible mutations, and in many cases the proteins survive and can fulfill some of their function. At the same time, some important changes can happen. For example, the protein can became permanently active, and such activity would lead to overgrowth of cells and eventually to cancer.”
Tsigelny added: “The clinical treatments that do not take into consideration the specific mutations in genes, but rather treat all mutations within a specific gene such as EGFR as the same, are a ‘hit-or-miss’ game because they are based solely on observational data on how other patients previously have reacted to drug therapies.”
This study is also important because it explains why some new mutations in EGFR – which are acquired during the course of treatment of lung cancer with small molecule EGFR inhibitors such as erlotinib – lead to drug resistance and patient death.
“The body doesn’t recognize tumors as harmful, but instead treats them like normal organs that need protection,” said Tsigelny. “This drives the development of mutations that cause drug resistance.”
Ready-Made Treatments
Moreover, new drugs are being developed to treat the resistant mutations, but the specific ones studied in this paper have not been amenable to therapy with these new drugs. Instead, the study showed that an existing drug (an antibody that targets EGFR) could be used, indicating that there may be a ready-made treatment available.
The modeling predictions are consistent with the clinical results. Two patients with the specific resistance mutation analyzed achieved partial remissions when treated with a regimen that included the EGFR antibody cetuximab, as predicted by the computer-based structural modeling. One patient continues to do well more than 3.5 years later.
“These studies form a proof-of-principle demonstration of the ability of computer modeling to be used to choose therapy for individuals,” said Razelle Kurzrock, senior deputy director (clinical science) and director of the Center for Personalized Cancer Therapy at UCSD Moores Cancer Center, and senior author of this paper. “Further investigation in larger patient groups is needed.”
Both Tsigelny and Kurzrock agreed that this finding is an excellent example of the power of collaboration between SDSC and the Moores Cancer Center, and that such modeling needed to be studied across tumors and with multiple different genes involved in cancer.
In addition to Tsigelny and Kurzrock, researchers involved in the study include Valentina L. Kouznetsova from SDSC and the Moores Cancer Center; Jerry P. Greenberg from SDSC; Lyudmila Bazhenova from the Moores Cancer Center; Jennifer J. Wheler from M.D. Anderson Cancer Center; and David J. Stewart from the University of Ottawa.
Financial support for the study was provided in part by the Joan and Irwin Jacobs Fund.
Share This:
You May Also Like
Stay in the Know
Keep up with all the latest from UC San Diego. Subscribe to the newsletter today.



