SDSC Researcher Awarded $1.4 Million NIH Structural Bioinformatics Grant
Three-year Grant to Increase Researcher Access to Archived Molecule Structures
Published Date
Article Content
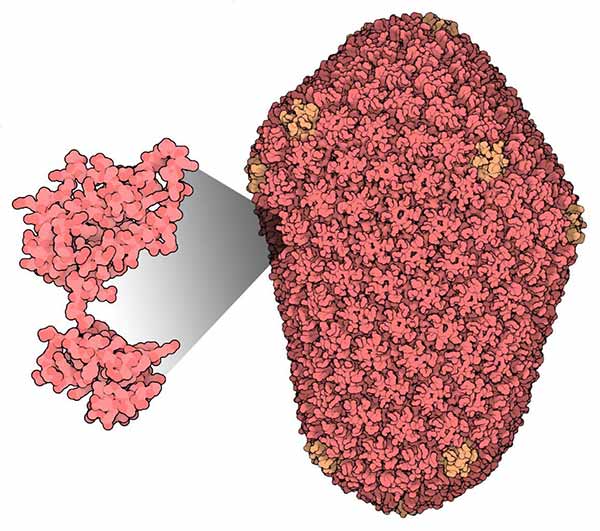
Structure of HIV Capsid (Source: RCSB Protein Data Bank)
A bioinformatics researcher at the San Diego Supercomputer Center (SDSC) at the University of California, San Diego, has been awarded a three-year National Institutes of Health (NIH) grant worth almost $1.4 million to make biological structures more widely available to scientists, educators, and students.
The NIH award, as part of the agency’s Targeted Software Development Awards and its Big Data to Knowledge (BD2K) initiative launched in 2012, was granted to Peter Rose, Site Head of the RCSB Protein Data Bank (PDB) West at SDSC and Project Scientist of the Center’s Structural Bioinformatics Laboratory, and Andreas Prlić, Technical and Scientific Team lead at the RCSB PDB.
The RCSB Protein Data Bank is the single worldwide repository for the three-dimensional structures of large molecules and nucleic acids that are vital to pharmacology and bioinformatics research. In May 2014, the PDB archived its 100,000th molecule structure, doubling its size in just six years. The 3-D structures, shapes of proteins and nucleic acids, now number about 111,000 and as the building blocks of life are fundamental to the understanding of disease processes, the mechanism of drug actions, and the development of new medicines.
“Currently, interactive visualization of large complex structures and structural comparisons across the entire Protein Data Bank archive exceeds available network bandwidth and the memory of typical scientists' desktops, laptops, or mobile devices,” said Rose, whose research interests also include large-scale “Big Data” mining, machine learning, and visualization of 3D structures, and its application in structure-based drug design. “This currently requires dedicated local network and a high-performance computing infrastructure that is not widely available.”
Part of the challenge is that the size and complexity of the structures have increased dramatically. For example, the recently determined structure of the HIV-capsid contains about 2.5 million atoms.
“In short, interactive visualization of large-scale structural analyses and queries of the archive have become a ‘Big Data’ or data-intensive challenge,” said Rose.
The project aims to make these structures accessible to a wider populace of researchers and students by developing a set of compression algorithms, applications, and workflows that will significantly improve the performance of interactive visualization of three-dimensional structures of large complexes over the Internet. Specifically, the goals are to:
- Develop a compact and extensible representation of 3-D biomolecular structures
- Enable interactive visualization of large complexes by reducing network bandwidth and enabling data streaming
- Enable large-scale analyses of the PDB archive for I/O bound workflows
- Develop open source software libraries.
“Such tools would enable users to analyze large structures, carry out large-scale searches, and visualize the coordinates directly in the compressed format,” said Rose. “There is great potential for discovery and innovation as the quantity and accessibility of biomedical data continues to expand. However, this potential can never be realized without the appropriate tools.”
“The RCSB PDB website is being used by 300,000 unique visitors per month,” said Prlić. “As such, the tools that are being developed as part of this grant will directly benefit a large community of scientists and data analysts. We are developing algorithms for the systematic comparison and analysis of all known protein structures. The data representations developed here will allow us to speed-up calculations significantly and will allow us to develop new algorithms for the analysis of large molecules. These are a big challenge for current approaches.”
The NIH award, called ‘Compressive Structural Bioinformatics: High Efficiency 3D Structure Compression’, is funded under grant number 1 U01 CA198942-01. The award is part of a series of BD2K awards announced for 2015.
Share This:
Stay in the Know
Keep up with all the latest from UC San Diego. Subscribe to the newsletter today.