Researchers Identify Elusive Carbon Dioxide Sensor in Plants that Controls Water Loss
Surprised biologists discover how two proteins work together to form long-sought plant water loss-regulating sensor, carrying implications for trees, crops and wildfires
Story by:
Published Date
Article Content
More than 50 years, ago researchers discovered that plants can sense carbon dioxide (CO2) concentrations. As CO2 levels change, “breathing” pores in leaves called stomata open and close, thus controlling evaporation of water, photosynthesis and plant growth. Plants lose more than 90% of their water by evaporation through stomata. The regulation of stomatal pore openings by CO2 is crucial for determining how much water plants lose, and is critical due to increased carbon dioxide effects on climate and water resources in a warming world.
But identifying the carbon dioxide sensor and explaining how it operates within plants has remained a longstanding puzzle.
UC San Diego biologists have unlocked the puzzle of how plants sense carbon dioxide, a key function as plants respond to climate in a warming world. iStock.com/mariaflaya
Using a mix of tools and research approaches, scientists at the University of California San Diego recently achieved a breakthrough in identifying the long-sought CO2 sensor in Arabidopsis plants and unraveled its functioning parts. UC San Diego project scientist Yohei Takahashi, School of Biological Sciences Distinguished Professor Julian Schroeder and their colleagues identified the CO2 sensor mechanism and detailed its genetic, biochemical, physiological and predicted structural properties. Their results are published December 7 in Science Advances.
Since the stomatal pores control plant water loss, the sensor is vital for water management and holds implications for climate-induced drought, wildfires and agricultural crop management.
“For each carbon dioxide molecule taken in, a typical plant loses some 200 to 500 water molecules to evaporation through the stomatal pores,” said Schroeder, Novartis Chair and faculty member in the Department of Cell and Developmental Biology. “The sensor is extremely relevant because it recognizes when CO2 concentrations go up and determines how much water a plant loses as carbon dioxide is taken in.”
One critical surprise from the new research was the composition of the sensor. Rather than tracing it to a single source or protein, the researchers found that the sensor operates through two plant proteins working together. These were identified as 1) a “high leaf temperature1” protein kinase known as HT1 and 2) specific members of a mitogen-activated protein kinase family, or “MAP” kinase enzyme, known as MPK4 and MPK12.
“Our findings reveal that plants sense changes in CO2 concentration by the reversible interaction of two proteins to regulate stomatal movements,” said Takahashi, who is now based at the Institute of Transformative Bio-Molecules, Japan. “This could provide us a new plant engineering and chemical target towards efficient plant water use and CO2 uptake from the atmosphere.”
The team’s findings, which have been filed in a UC San Diego patent, could lead to innovations in efficient water use by plants as CO2 levels rise.
“This finding is relevant for crops but also for trees and their deep roots that can dry out soils if there’s no rain for long periods, which can lead to wildfires,” said Schroeder. “If we can use this new information to help trees respond better to increases in CO2 in the atmosphere, it’s possible they would more slowly dry out the soil. Similarly, the water use efficiency of crops could be improved—more crop per drop.”
To further explore their sensor discovery, the researchers collaborated with graduate student Christian Seitz and Professor Andrew McCammon in the Department of Chemistry and Biochemistry. Using cutting-edge techniques, Seitz and McCammon created a detailed model of the intricate structure of the sensor. The model implicated areas where genetic mutations have been known to restrict the ability of plants to regulate transpiration in response to carbon dioxide. The new imagery showed that the mutants cluster in an area where the two sensor proteins, HT1 and MPK, come together.
“If we can use this new information to help trees respond better to increases in CO2 in the atmosphere, it’s possible they would more slowly dry out the soil. Similarly, the water use efficiency of crops could be improved—more crop per drop.”
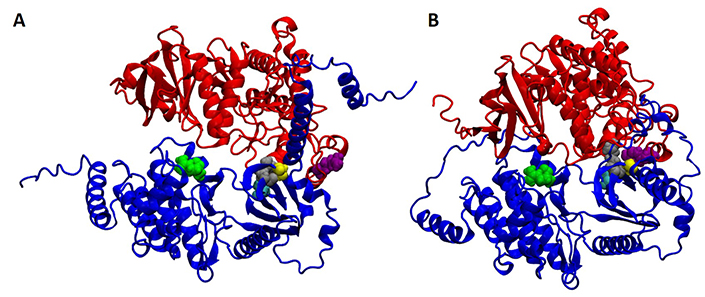
Working with colleagues in the Department of Chemistry and Biochemistry, UC San Diego biologists unraveled the predicted structure of the newly discovered plant carbon dioxide sensor. The left section (A) depicts the MPK4 – HT1 complex (MPK highlighted in red; HT1 in blue) and the right (B) section reveals the MPK12 – HT1 complex. The highlighted amino acid residues (yellow, gray, light blue and green) show mutations that disrupt the sensor function. Credit: McCammon Lab, UC San Diego
“This work is a wonderful example of curiosity-driven research that brings together several disciplines—from genetics to modeling to systems biology—and results in new knowledge with the ability to aid society, in this case by making more robust crops,” said Matthew Buechner, a program director in the U.S. National Science Foundation’s Directorate for Biological Sciences, which supported the research.
The paper’s full author list: Yohei Takahashi, Krystal Bosmans, Po-Kai Hsu, Karnelia Paul, Christian Seitz, Chung-Yueh Yeh, Yuh-Shuh Wang, Dmitry Yarmolinsky, Maija Sierla, Triin Vahisalu, J. Andrew McCammon, Jaakko Kangasjarvi, Li Zhang, Hannes Kollist, Thien Trac and Julian I. Schroeder.
Funding for the research described in the Science Advances paper was provided by the NSF (grant MCB-1900567); and in part by the National Institutes of Health (grant R01 GM60396); a National Science Foundation Graduate Research Fellowship (DGE-1650112), JST, PRESTO (grant JPMJPR21D8); SUNBOR; the Estonian Research Council (grant PRG433); Centre of Excellence CEMCE; Plant Biology Infrastructure project TAIM; and the Academy of Finland Centre of Excellence program (grants 271832 and 307335).
Learn more about research and education at UC San Diego in: Climate Change
Share This:
You May Also Like
UC San Diego is Strengthening U.S. Semiconductor Innovation and Workforce Development
Technology & EngineeringStay in the Know
Keep up with all the latest from UC San Diego. Subscribe to the newsletter today.



