Researchers Discover Hidden SARS-CoV-2 ‘Gate’ That Opens to Allow COVID Infection
Supercomputing-derived movies reveal details of deceptive sugar coating on spike protein, presenting new possibilities to block cell entry and infection
By:
- Mario Aguilera
Published Date
By:
- Mario Aguilera
Share This:
Article Content
The glycan gate opens: Supercomputing-driven simulations depict the glycan N343 (magenta) acting as a molecular crowbar to pry open the SARS-CoV-2 spike’s receptor binding domain, or RBD (cyan), from a “down” to an "up" position. Credit: Terra Sztain, Surl-Hee Ahn, Lorenzo Casalino (Amaro Lab, UC San Diego).
Since the early days of the COVID pandemic, scientists have aggressively pursued the secrets of the mechanisms that allow severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) to enter and infect healthy human cells.
Early in the pandemic, University of California San Diego’s Rommie Amaro, a computational biophysical chemist, helped develop a detailed visualization of the SARS-CoV-2 spike protein that efficiently latches onto our cell receptors.
Now, Amaro and her research colleagues from UC San Diego, University of Pittsburgh, University of Texas at Austin, Columbia University and University of Wisconsin-Milwaukee have discovered how glycans—molecules that make up a sugary residue around the edges of the spike protein—act as infection gateways.
Published August 19 in the journal Nature Chemistry, a research study led by Amaro, co-senior author Lillian Chong at the University of Pittsburgh, first author and UC San Diego graduate student Terra Sztain and co-first author and UC San Diego postdoctoral scholar Surl-Hee Ahn, describes the discovery of glycan “gates” that open to allow SARS-CoV-2 entry.
“We essentially figured out how the spike actually opens and infects,” said Amaro, a professor of chemistry and biochemistry and a senior author of the new study. “We’ve unlocked an important secret of the spike in how it infects cells. Without this gate the virus basically is rendered incapable of infection.”

Rommie Amaro
Amaro believes the research team’s gate discovery opens potential avenues for new therapeutics to counter SARS-CoV-2 infection. If glycan gates could be pharmacologically locked in the closed position, then the virus is effectively prevented from opening to entry and infection.
The spike’s coating of glycans helps deceive the human immune system since it comes across as nothing more than a sugary residue. Previous technologies that imaged these structures depicted glycans in static open or closed positions, which initially didn’t draw much interest from scientists. Supercomputing simulations then allowed the researchers to develop dynamic movies that revealed glycan gates activating from one position to another, offering an unprecedented piece of the infection story.
“We were actually able to watch the opening and closing,” said Amaro. “That’s one of the really cool things these simulations give you—the ability to see really detailed movies. When you watch them you realize you’re seeing something that we otherwise would have ignored. You look at just the closed structure, and then you look at the open structure, and it doesn’t look like anything special. It’s only because we captured the movie of the whole process that you actually see it doing its thing.”
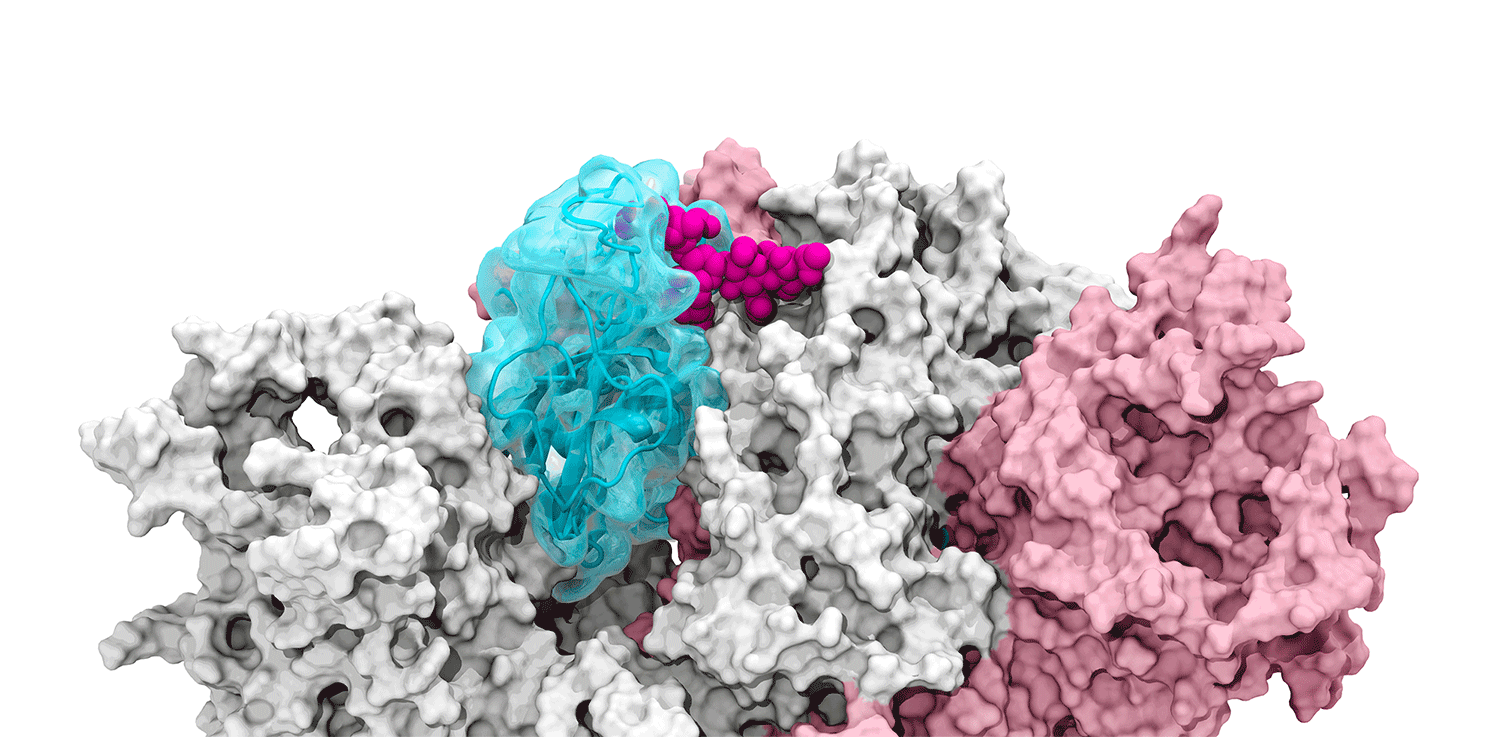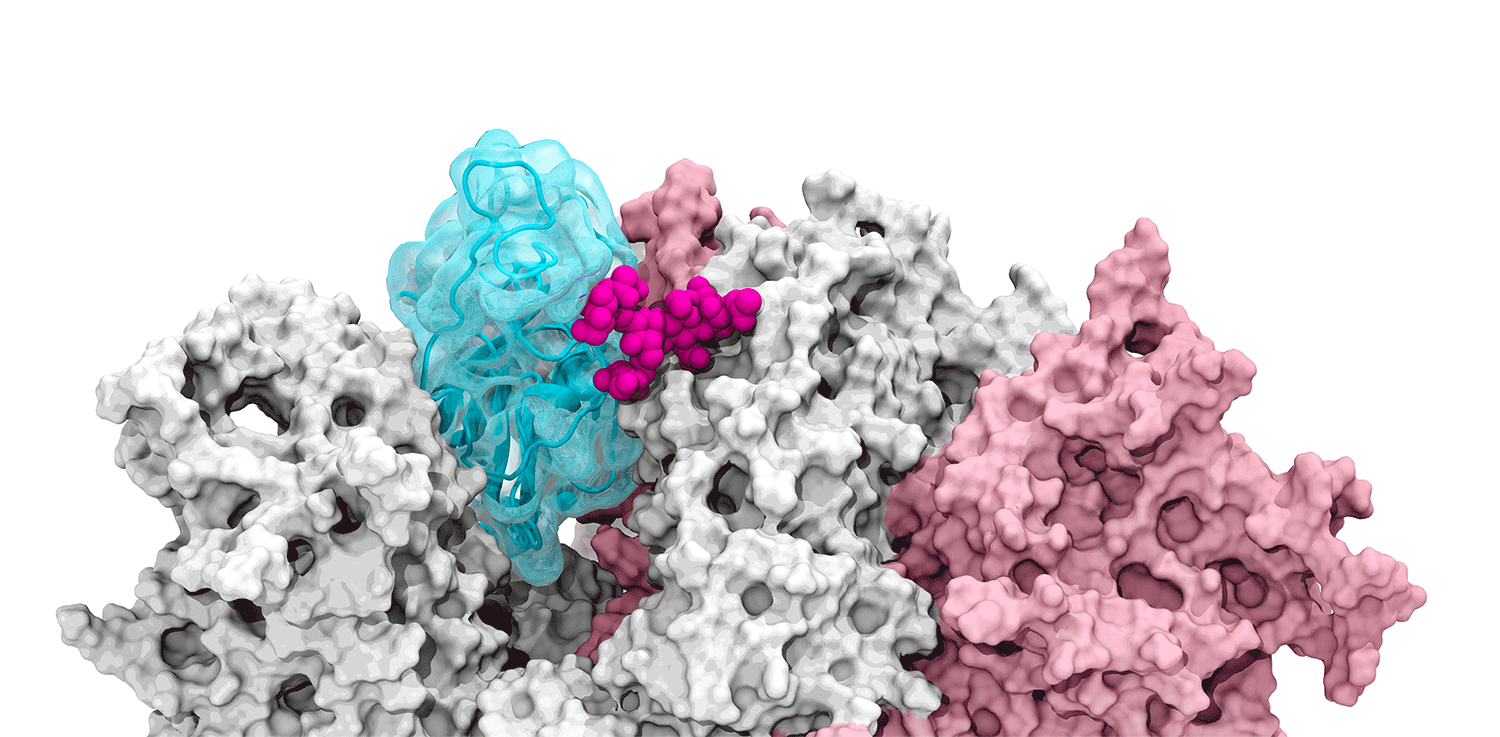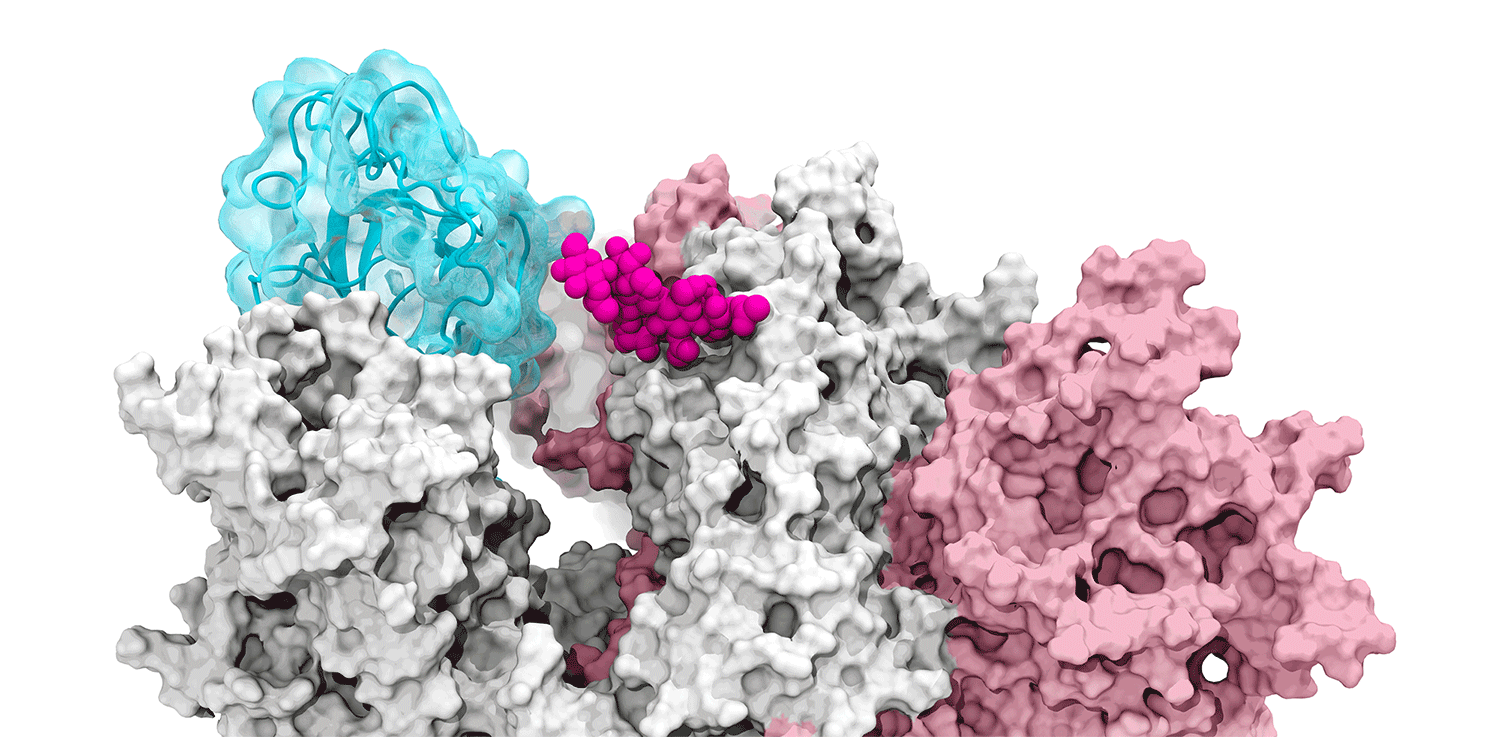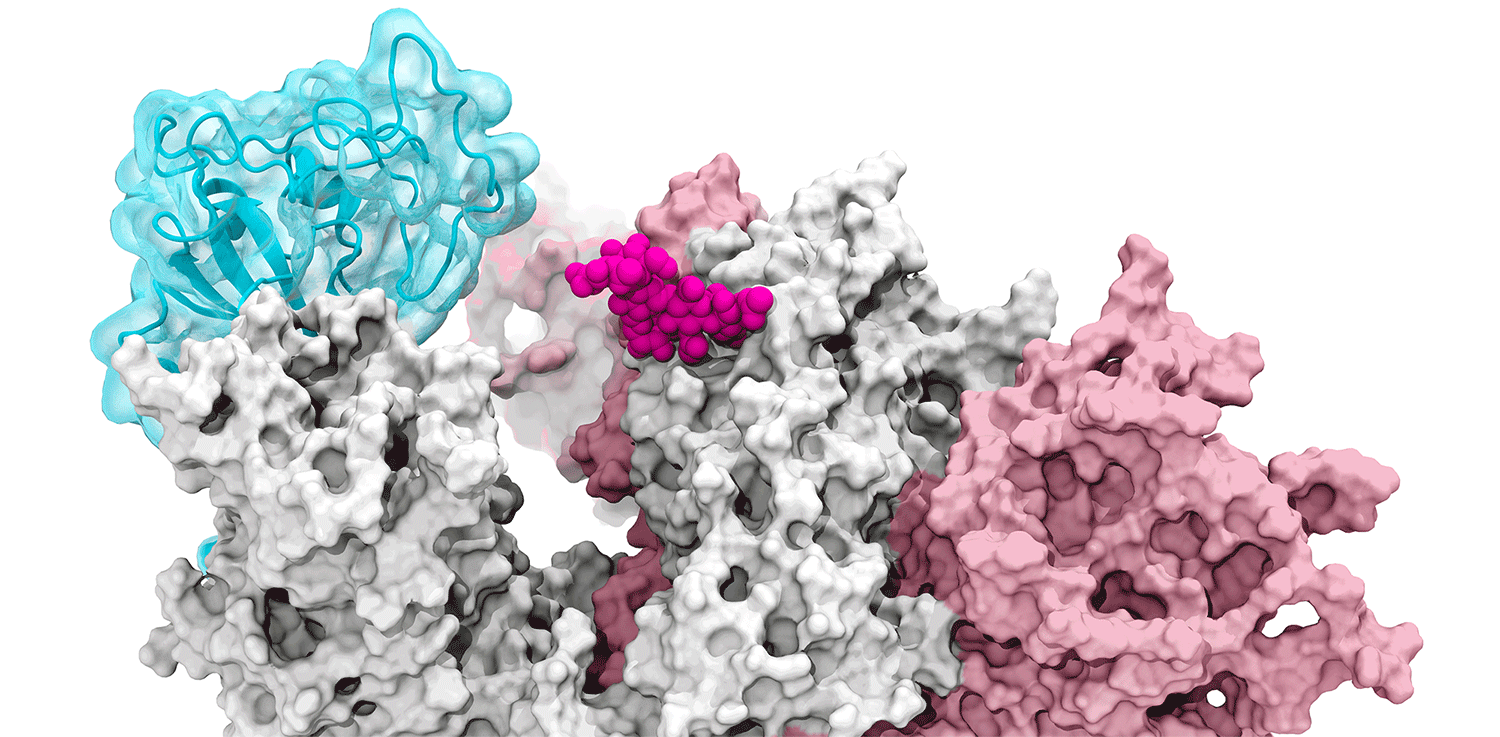
The glycan gate opens: Supercomputing-driven simulations depict the glycan N343 (magenta) acting as a molecular crowbar to pry open the SARS-CoV-2 spike’s receptor binding domain, or RBD (cyan), from a “down” to an "up" position. Credit: Terra Sztain, Surl-Hee Ahn, Lorenzo Casalino (Amaro Lab, UC San Diego).
“Standard techniques would have required years to simulate this opening process, but with my lab’s ‘weighted ensemble’ advanced simulation tools, we were able to capture the process in only 45 days,” said Chong.
The computationally intensive simulations were first run on Comet at the San Diego Supercomputer Center at UC San Diego and later on Longhorn at the Texas Advanced Computing Center (TACC) at UT Austin. Such computing power provided the researchers with atomic-level views of the spike protein receptor binding domain, or RBD, from more than 300 perspectives. The investigations revealed glycan “N343” as the linchpin that pries the RBD from the “down” to “up” position to allow access to the host cell’s ACE2 receptor. The researchers describe N343 glycan activation as similar to a “molecular crowbar” mechanism.
Jason McLellan, an associate professor of molecular biosciences at UT Austin and his team created variants of the spike protein and tested to see how a lack of the glycan gate affected the RBD’s ability to open.
“We showed that without this gate, the RBD of the spike protein can’t take the conformation it needs to infect cells,” McLellan said.
“TACC has been essential for our work since 2004, through allocations on XSEDE for molecular dynamics research projects on bioengineering and cellular biology,” said Amaro. “For COVID-19, Frontera and Longhorn have been absolutely instrumental for our all-atom simulations of the SARS-CoV-2 spike protein leading up to the whole virion.”
The full author list includes: Terra Sztain, Surl-Hee Ahn, Anthony Bogetti, Lorenzo Casalino, Jory Goldsmith, Evan Seitz, Ryan McCool, Fiona Kearns, Francisco Acosta-Reyes, Suvrajit Maji, Ghoncheh Mashayekhi, J. Andrew McCammon, Abbas Ourmazd, Joachim Frank, Jason McLellan, Lillian Chong and Rommie Amaro.
The research was funded by: the National Science Foundation (NSF) (GRFP DGE-1650112); the National Institutes of Health (NIH) (GM132826); NSF (RAPID MCB-2032054); the RCSA Research Corp.; a UC San Diego Moores Cancer Center 2020 SARS-COV-2 seed grant; NIH (R01-GM31749); NIH (R01-AI127521); NIH (R01 GM115805); NSF (CHE-1807301); and National Institute of General Medical Science (R01 GM29169 and R35 GM139453).
Share This:
You May Also Like
Stay in the Know
Keep up with all the latest from UC San Diego. Subscribe to the newsletter today.



