Leading-edge Technology Unmasks Protein Linked to Parkinson’s Disease
Two cryo-EM studies pave the way for possible drugs to treat neurodegenerative disorder affecting millions
Published Date
Article Content
An elusive protein that many consider the key of fully understanding the causes of genetic Parkinson’s disease has come much more clearly into focus.
Impacting millions around the world, Parkinson’s is a neurological disorder that progressively attacks motor functions, leading to lasting damage in movement and coordination, among other areas.
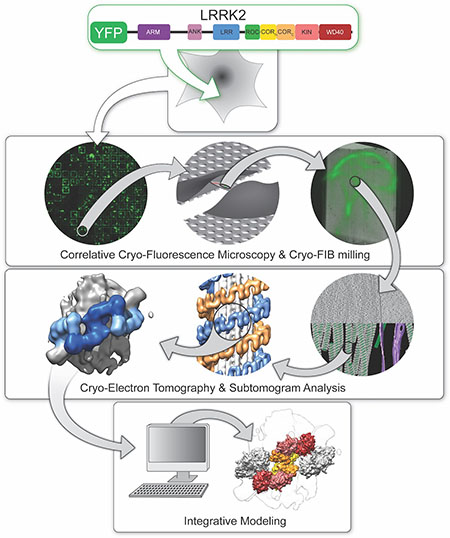
State-of-the-art technologies researchers used to finally unveil the structure of the LRRK2 enzyme inside cells. First, they used fluorescence proteins to find it, then they used a beam of ions to carve out regions that contained LRRK2 filaments, which they imaged using cryo-electron tomography. Finally, they computationally analyzed the data to build models of the protein in high level of detail.
Researchers studying the primary causes of the disease have focused on mutations of the protein known as leucine-rich repeat kinase 2, or LRRK2. Understanding how LRRK2 disrupts normal functioning has been difficult due to a lack of information on the structure of the protein. Since LRRK2 is a major drug target, efforts to decipher LRRK2’s architecture have even included launching samples into space as a way of using microgravity conditions to help crystalize protein samples, but with no success.
Now, University of California San Diego scientists using leading-edge technologies have produced the first visualizations of LRRK2 inside its natural cellular environment and the first high-resolution blueprint of the protein. They leveraged these depictions to describe how LRRK2 binds to cellular tracks called microtubules and acts as a roadblock for motors that move along these tracks. The findings are described in two research papers published in the journals Cell and Nature.
"These two papers take giant steps towards developing more effective therapeutics for Parkinson’s Disease, which impacts so many lives,” said Division of Biological Sciences Dean Kit Pogliano, PhD, a professor in the Section of Molecular Biology. “Combining cryo-electron microscopy with live cell imaging allows researchers to see proteins working inside cells and to more rapidly determine how potential drugs affect their function. This will accelerate drug discovery and provide new hope to those suffering from this debilitating disease.
LRRK2: a key therapeutic target for Parkinson’s disease
Known as a kinase, LRRK2 is a type of enzyme that adds phosphate chemical tags to other proteins, influencing their function. LRRK2 mutations are the main cause of inherited forms of Parkinson’s disease, but researchers do not fully understand how this enzyme functions normally or in disease states.
The answers to such questions have been vigorously pursued, especially because kinases are one of the most established pharmacological targets for therapeutics. Since the LRRK2 gene was first cloned in 2004, there has been a tremendous amount of effort devoted to targeting it for Parkinson’s treatment.
Susan Taylor, a world-renowned authority in kinases, recognized that the expertise at UC San Diego was ideally suited to tackle this problem. With support from the Michael J. Fox Foundation https://www.michaeljfox.org, an international team, including researchers across UC San Diego, set out to study LRRK2 using novel technologies.
A cellular blueprint of LRRK2 on microtubules
As described in the Cell paper, UC San Diego researchers led by Elizabeth Villa, PhD, and her colleagues leveraged new groundbreaking technology known as cryo-electron tomography (cryo-ET), a modality of cryo-electron microscopy (cryo-EM), to view LRRK2 in its natural environment within cells and describe its structure at a level previously unseen.
In many cases, when researchers seek to determine a protein’s structure, they begin by isolating the protein outside of cells. When using cryo-EM, scientists freeze the molecules in a thin layer of ice, preserving their structure, and determine their structure at high resolution. Instead, Villa’s team images frozen cells that contain the molecules under study, taking pictures at different angles, somewhat like a CAT scan.
“We lifted the hood to look inside the cell at frozen molecules caught in the act of interacting with each other in different places in the cell,” said Villa, who noted the significance of co-developing the cutting-edge instruments used in the study with Thermo Fisher Scientific and UC San Diego’s considerable efforts in establishing a new cryo-EM facility on campus. “We used electron and ion beams like a light saber to blast away parts of the cell. In the middle we left a window that contains the molecules that we are interested in looking at.”
The researchers also used light microscopy to find the molecules within cells, and advanced computational modeling tools to develop a high-resolution integrative model of a mutant LRRK2. Their data revealed LRRK2 bound to cellular highways, called microtubules, decorating them in a stunningly regular and unexpected geometry, and predicted that LRRK2’s kinase resembled an “overdrive” state known to occur in Parkinson’s disease.
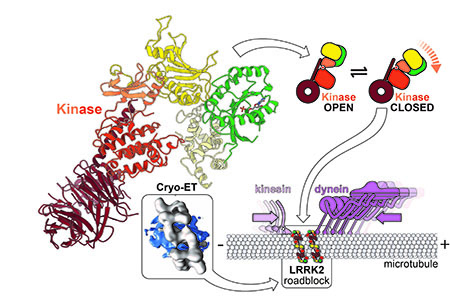
UC San Diego researchers discovered that LRRK2 binds microtubules and creates roadblocks that stop the molecular machines that drive cellular cargo transport. They also showed that some drugs that target LRRK2’s kinase enhance this effect, while others diminish it.
“This combination of powerful new techniques was applied for the first time in this research and made it possible to get the first glimpses into the structure of a mutant LRRK2, with the added benefit of its cellular context,” said Villa. “To our knowledge it’s the highest resolution structure of a human protein that was determined inside cells before using conventional biochemical tools. We are bringing structure to cell biology.”
A molecular blueprint of LRRK2
In order to understand how LRRK2 works at a chemical level and to design therapeutics, an even higher resolution structure is required to reveal the position of atoms and how they interact with potential drugs. In the Nature study, co-senior authors Samara Reck-Peterson, PhD, and Andres Leschziner, PhD, took a deeper look at LRRK2’s structure and function and teamed up with Villa’s group to determine how LRRK2 interacts with microtubules.
Using cryo-EM, Leschziner’s team captured the most detailed image-to-date of LRRK2’s structure, down to the atomic level. Stefan Knapp, a professor at Goethe University in Frankfurt Germany, and his group were also instrumental as they determined how to make LRRK2 tractable for structural work. The structure comprised the business end of the protein — which includes the part that tags other proteins with phosphates. The locations of all major Parkinson’s disease-causing mutations are found in this structure.
Next, the Leschziner group combined their structure with Villa’s and came up with a model that explains how LRRK2 binds to microtubules.
“You can think of the kinase part of LRRK2 as a bit like Pacman, it can be either open or closed,” said Leschziner, professor at UC San Diego School of Medicine and Division of Biological Sciences. “Our modeling suggested that, when bound to microtubules, the kinase needs to be in a closed state, indicating that the shape of the kinase may regulate the binding of LRRK2 to microtubules. Teaming up with the Reck-Peterson lab we decided to test this model directly.”
LRRK2 is a roadblock for molecular motors
Microtubules are Reck-Peterson’s specialty. She and her team are interested in the molecular motors that transport cargo along microtubules, and how defects in this transport cause human neurodevelopmental and neurodegenerative diseases. Reck-Peterson and her team wondered if LRRK2’s interaction with microtubules might be detrimental for the molecular machines that move on them and carry essential cargos from place to place in cells.
Her team discovered that LRRK2 creates roadblocks that stop these molecular machines. They also showed that some drugs that target LRRK2’s kinase enhance this effect, while others diminish it. While Leschziner and Reck-Peterson are not sure yet if roadblocks play a role in Parkinson’s disease, their findings already have implications for the design of therapeutic drugs that work by inhibiting LRRK2. Their work points to the possibility that kinase inhibitors that close LRRK2’s kinase domain might lead to the unwanted effect of blocking the movement of molecular motors.
“It’s not yet clear what role LRRK2-microtubule binding plays in Parkinson’s disease,” said Reck-Peterson, professor at UC San Diego School of Medicine and Division of Biological Sciences and a Howard Hughes Medical Institute investigator. “But what we have now are cellular and molecular blueprints, and that’s what is needed to figure out what LRRK2 does and to fine-tune therapeutic drugs that target LRRK2.”
Co-authors of the Cell study: Reika Watanabe, Robert Buschauer, Jan Böhning, Martina Audagnotto, Tsan Wen Lu, Daniela Boassa, Susan S. Taylor and Elizabeth Villa, UC San Diego; Keren Lasker, Stanford University.
The Cell study was funded by: National Institutes of Health Director’s New Innovator Award (1DP2GM123494-01), a Michael J. Fox Foundation grant (11425), an NIH National Institute of General Medical Science (R01GM086197), a Branfman Family Foundation award, a postdoctoral fellowship from the Visible Molecular Cell Consortium at UC San Diego and a Taiwan MOE fellowship.
Co-authors of the Nature study: Colin K. Deniston, David M. Snead, Indarjit Lahiri, Reika Watanabe, Jan Böhning, Elizabeth Villa, Samara L. Reck-Peterson, and Andres E. Leschziner, UC San Diego; John Salogiannis, Oscar Donosa, Howard Hughes Medical Institute; Sebastian Mathea, Stefan Knapp, Goethe-Universitat; Andrew K. Shiau, Ludwig Institute for Cancer Research
The Nature study was funded by: the National Institutes of Health (grants 1S10RR23057, 1S10OD018111, 1U24GM116792, T32 GM008326, DP2GM123494, R01GM121772, R01GM107214), National Science Foundation (grant DBI-1338135), Michael J Fox Foundation (grants 11425 and 11425.02), CNSI at UCLA, UC San Diego Cryo-EM Facility, UC San Diego Visible Molecular Cell Consortium and Center for Trans-scale Structural Biology, A.P. Giannini Foundation, Ludwig Institute for Cancer Research, Howard Hughes Medical Institute, DFG (grant HE1818/11), Boehringer Ingelheim, and SGC, a registered charity that receives funds from AbbVie, Bayer Pharma AG, Boehringer Ingelheim, Canada Foundation for Innovation, Eshelman Institute for Innovation, Genome Canada, Innovative Medicines Initiative [ULTRA-DD], Janssen, Merck, KGaA Darmstadt Germany, MSD, Novartis Pharma AG, Ontario Ministry of Economic Development and Innovation, Pfizer, São Paulo Research Foundation-FAPESP, Takeda and the Wellcome Trust.
Share This:
Stay in the Know
Keep up with all the latest from UC San Diego. Subscribe to the newsletter today.